Goals
- Outline the process of translation
- Describe the structure and role of tRNAs
- Explain and contrast the process of translation initiation in eukaryotes and prokaryotes
- Describe the role of the A,P, and E sites in the ribosome during translation elongation
- Explain how release factor terminates translation
- Explain how the relationship between transcription and translation differs between eukaryotes and prokaryotes
Overview of Translation
The process of translation requires an enzyme: Ribosome; a template: mRNA; subunits: amino acids; an intermediary molecule; tRNA’s
The key steps of translation are: initiation, elongation, and termination
Ribosome
A ribosome, also known as a ribonulceoprotein complex consists of 3 RNA’s and 50+ proteins. There is a small and large subunit.
Major drug target (antibiotics)
tRNA
tRNA’s work by bringing the specific amino acid to the ribosome where translation is occurring. The tRNA is the bridge between the mRNA and the amino acid.
Every amino acid delivered has a a different tRNA, and are recognized by the codon by the anticodon on the tRNA which is complementary to the tRNA
mRNA
Structure
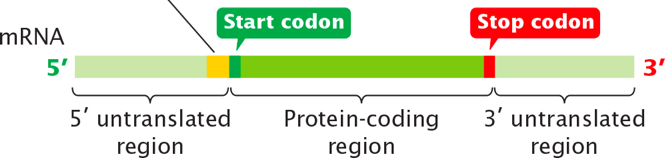
mRNA is the template for translation. At the 5′ end has an untranslated region upstream of the AUG (start codon), and at the 3′ end also has an untranslated region downstream of the stop codon.
It has an initiation factor called the start codon, and a stop codon to terminate translation. In between those two regions there is the protein coding-region where the sequences in this region are translated into a poly-amino acid strand.
Translation Initiation
Prokaryotes
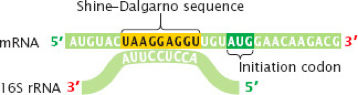
The binding site of the ribosome is a specific seqeunce in the mRNA which is known as Shine-Dalgarno sequence, and occurs three base pairs upstream of the initiation codon.
This sequence is seen above, and is UAAGGAGGU, and the Shine-Dalgarno sequence is recognized by the ribsome complementary base pairing with the mRNA at this location.
The small ribosomal subunit and an initiator tRNA is first to bind at the Shine-Dalgarno sequence or in other words the ribosome binding site, and it is followed by the addition of the by the large ribosomal subunit, and initiation commences.
Prokaryotes are able to have many independent ribosomal binding sites, for different independent proteins those biding sites are assigned to in the transcript.
Eukaryotes
Transcription begins by the addition of a diverse set of factors known as eukaryotic initiation factors or (eIFs) which bind the mRNA. They ultimately “load on” the small ribosomal subunit (40S)
The small ribosomal subunit then goes on to “scan” the mRNA to detect the translation initiation site (AUG), and once it is found the large ribosomal subunit proceeds to join the small subunit and initiates translation.
Eukaryotic mRNA can also become circularized to promote the re-initiation of translation to make duplicates/ more copies of the protein.
Translation Elongation
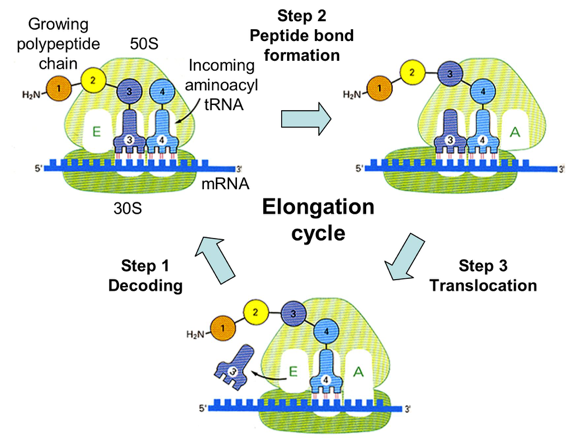
There are three sites on the ribsome which are significant to translational elongation. There is the A site, the P site, and the E site.
The A site: Aminoacyl tRNA site
The Aminoacyl tRNA site is the site where the next tRNA’s, containing the next Amino Acid that is to be added, anticodon complimentary base pairs to the codon of the mRNA strand.
The P Site: Peptidyl tRNA site
The Peptidyl tRNA site is the site where the tRNA from the previous A-site, after moving over now to the P-site, is bound to the existing chain by the ribosome.
The E Site: Exit Site
Just like the name says, this is the exit site. tRNA leaves the ribosome after the amino acid it once carried was bound to the existing protein chain, and has shifted towards the E site.
rRNA
The Ribosomes rRNA is thought to be the ribozyme catalyst, with evidence shown when the 50S subunit is treated with proteinase K, which degrades proteins), but is still capable of catalytic activity. Structurally the proteins are not near the active site.
Polyribosomes
mRNA is translated by many ribosomes at the same time.
Translation Termination
When the stop codon on the mRNA is at the A site a release factor protein (which structurally resembles a tRNA) binds to the stop codon which leads the commences termination. This causes hydrolysis of the peptide chain from the P site tRNA, and leads to dissociation of the ribosome ,mRNA, and tRNAs
Coupling of Transcription and Translation
In prokaryotes, translation and transcription are coupled.
In eukaryotes, transcription and translation are separate in timing and location.
Leave a comment