Goals
- Describe the structure and function of the 5′ cap and polyA tail of an mRNA
- Explain the mechanism of splicing and how it is enhanced by the spliceosome
- Explain how and why transcription and mRNA processing are coupled
- Draw an RNApolII transcript including relevant structural features (cap, polyA tail, introns, exons)
- Explain the role of mRNA processing in nuclear export
Eukaryotic Transcription: mRNA Production.
mRNA processing proteins bind to elongating RNA PolII, and once the protein elongates the mRNA, the first processing, called a capping factor, protein binds to the 5′ end of the mRNA. This is known as the 5′ cap.
This protein capping process places a 5′ 7-methylguanosine cap (m7G) creating a 5′-5′ phosphate linkage between the mRNA strand, and the m7G cap known as a 5′ to 5′ triphosphate bridge. This protects the mRNA from degradation, and can also be bound by cap specific proteins
Eukaryotic Transcription: Termination
In Eukaryotic transcription, the sequence AAUAAA is recognized in the end of the mRNA transcript (known as the poly-A signal), which signals the end of the message. This signal releases the mRNA transcript, and then as a result a Rat1/hXm2 RNA cleaves the mRNA from RNA Polymerase II (RNApolII), and the transcript is released.
Immediately upon release, PolyA polymerase adds the polyA tails at the poly-A signal sequence immediately post release. This protects the mRNA at the 3′ end, and is also bound by the polyA specific proteins.
- The PolyA tail is added by PolyA Polymerase (PAP), which adds ~100-250 nucleotides at the 3′ end of the mRNA
RNA polymerase is still elongating more mRNA along the strand, and the same nuclease protein Rat1/hXm2 starts a exonuclease event on the mRNA still being produced as it doesn’t have a protective cap.
RNA splicing
mRNA that has just been elongated, is actually known as pre-mRNA because it has not been spliced yet. The pre-mRNA has sequences known as exons and introns. The introns are spliced out of the mRNA, as they do not go on to become the mature mRNA. They are removed via splicing, and the remaining fragments called exons are religated to form the mature mRNA.
Introns are relatively larger than exons sizing about greater 100 nucleotides and to thousands of nucleotides. Exons however are much smaller ranging from 50 nucleotides (typically) to a few hundred nucleotides in size.
Introns are also not “junk” mRNA, but the may include regulatory sequences and/or small functional RNAs.
Since the names of introns and exons can be confusing when studying this subject, just remember that introns are removed because they are intervening species, and exons are ligated together because exons expressed. Introns are big, with small sequences inersepersed (exons) of the final product.
There is experimental evidence of introns as the experimenters mixed mature mRNA and DNA together, they made the DNA single stranded and the mRNA linear. Then they allowed them to anneals together where the DNA can complementary base pair to the RNA, forming a DNA/RNA hybrid. They were visualized by microscopy, and what they seen was with a gene that had no introns the strand of RNA was able to completely base pair. However when introns were present in the gene, and the mature mRNA re-annealed to the complementary DNA (cDNA) is visualized to be interspersed where the exons complementary base paired to their corresponding regions on the cDNA, and the DNA forms loops at the single stranded areas.
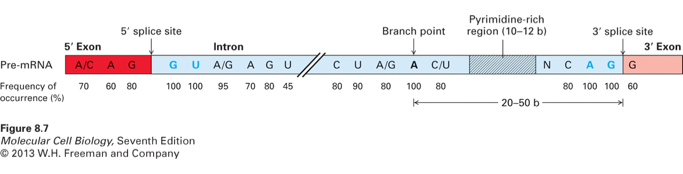
There are specific splice sights located on the pre-mRNA in the intron regions, 100% of the time at the 5′ splice there is a GU located in the intron. Downstream of the 5′ (GU) splice site, 95% of the time there is either an A or a G. In the middle of the intron, known as the branch point 100% of the time there is an A, and at the 3′ splice site in the intron 100% of the time there is an AG.
The reason why the exons do not contain the splice sites, is because they must have a specific sequence which continue on to produce proteins. The introns, however, do not need a specific sequence, thus, contains the splice site.
Near these splice sites percentages of specific nucleotides are high to indicate the splice site is near.
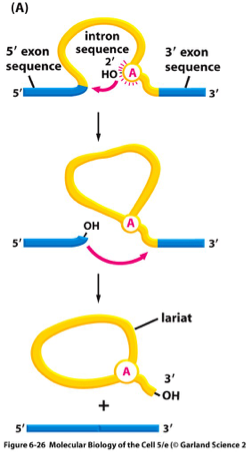
The introns are removed because the 3′ hydroxyl group (3′-OH) of the A at the branch point attacks the G at the 5′ splice sites. The the free OH group on the 3′ exon then attacks the G on the 3′ splice site, and ultimately sees the intron exiting as a lariat structure, and the two exons are ligated together.
This process isn’t done by itself. There are proteins, that come to the rescue. These proteins combined are known as a the spliceosome. These proteins are involved in splicing. They are a complex of snRNA’s and proteins. Each 5 snRNAs (U1, U2, U6 & U4 complex, U5, and U6 U4 and U5 complex) joins with a protien to form an individual ribonucleoprotein complex known as a snRNP (pronounced a snurp), and all 5 snRNP’s together forms the spliceosome .
The snRNA’s complementary base pairs to the pre-mRNA, and are likely catalytic. The snRNP’s position the RNA for subunits for catalysis, and they remodel the spliceosome during splicing.
It is thought that snRNA’s are most likely catalytic, and a re therefore integral parts of the spliceosome. The U1 snRNP recognizes the 5′ (GU) splice site, the U2 snRNP recognizes the A branch point, and the U4/U6 snRNP and U5 snRNP join the previous two snRNP’s and the spliceosome is formed. Conformation of the intron is remodelled allowing for the attack of the branch point to the 5′ splice site. This is the first cleavage event.
After the first cleavage event, the intron is remodelled again, and promotes the second cleavage event and the joining of the exons.
pre-mRNA can also self splice, this occurs in mitochondria and chloroplasts in some fungi, plants, and algae. It may be the evolutionary precursor to the spliceosome.
Splicing is very important, as 15% of disease-related human point mutations create splicing defects, and many proteins activat and inhibit splicing at specific sites.
Nuclear Export
Processed mRNA is exported from nucleus to the cytoplasm, by a process not discussed here. The mature mRNA is processed in the nucleus.
Proteins dictate nuclear export by binding to the mRNA during processing. They ensure the mRNA has a cap, a polyA tail, and the mRNA has been spliced of its introns. These proteins thus filter the unprocessed pre-mRNA and not approved mRNA by checking the proteins (which are left by the splicesome acting as a marker)
Leave a comment